What Can Enterprises Learn From Genome Sequencing?
The data handing requirements for genetic discovery are increasing as the tools become more sophisticated, causing data sets to become increasingly large. However, dealing with these large data sets is nothing new for practitioners in traditional science, and enterprises can learn from the strategies and processes that these disciplines have put into play.
 Mario Caccamo, head of Bioinformatics at The Genome Analysis Center (TGAC) based in the UK, recently outlined the work and challenges that researchers face, explaining the explosive nature of the data and the processes that that genome scientists use to wrestle with it.
Mario Caccamo, head of Bioinformatics at The Genome Analysis Center (TGAC) based in the UK, recently outlined the work and challenges that researchers face, explaining the explosive nature of the data and the processes that that genome scientists use to wrestle with it.
“We have super exponential growth in data generation,” explained Caccamo. “To give you a concrete example, with two sequencing instruments in 2010, we generated 1.2 terabases of data. We can now generate half of that (600 gigabases) in two weeks in only one sequencing round with our current instruments.”
This, of course, is a familiar refrain that enterprise executives are hearing as they consider their own data plans and consider what their own data requirements will be down the line. But having an infrastructure to handle and store this data isn’t the same thing as having a strategy to turn it into predictive intelligence.
“This is very much hitting the point where you need a strategy to cope with more data today than you have ever generated before,” says Caccamo, explaining the growing challenges TGAC faces as their data volumes increase in size and complexity. The organization looks at these challenges holistically, explained Caccamo in describing how they have built a cultural infrastructure to support their data-driven science goals.
Caccamo explains that a technology focus drives their cultural underpinnings. “In one end, developing the state of the art intensive platform, both sequencing and computational – this is what I call the ‘hard infrastructure.’ The other focus of our activities is on what we can develop between intensive algorithms and databases – this is what we call the ‘soft infrastructure,’”
Caccamo explains that TGAC’s challenge is on developing these components together through what he calls a systems approach in order to produce “predictive assumptions” that help them towards discovery and ultimately understanding biology.
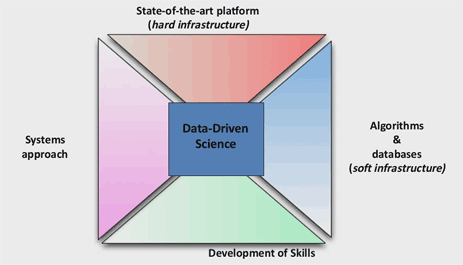
Included in their base equation is the development of skills, says Caccamo, being very sensitive to the talent acquisition challenges that organizations face in the exploding data science fields. “We really take training into this,” explained Caccamo. “Technologies and training toward developing new strategies and tools is a very important part of what we do.”
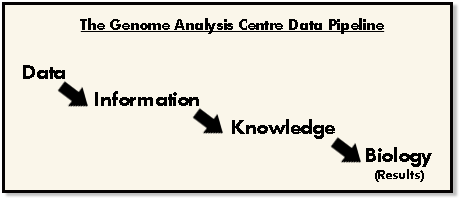
Putting this organizational foundation into place, explained Caccamo, they are then able to give focus to what their true aim is: data driven science. Their process is something that other organizations can learn from, suggests Caccamo, explaining TGAC’s view of the pipeline that starts with data on the input end, and biology science as the output.
The process starts with huge volumes of sequencing data and using efficient algorithms to tackle the data, endeavor to distill it into what can then be classified as “information.” In the case of genomic research, the information are the ~150 letter base strings that the researchers use to assemble genome maps with. The research head says that once they have achieved the information step in the process, the focus, they shift priorities from efficiency to quality. At this step, says Caccamo, they’re focused on turning “information” into “knowledge” which can then be transformed into biology.
In the case of genome sequencing, the research head used the discovery process of the wheat genome as an example. The first step is in extracting and sequencing the DNA into strands of information called “bases,” which are ready for assembling into something more actionable.
These bases are then passed through sophisticated hardware (TGAC uses an array of SGI UV supercomputers for much of their assembly work) and assembled into enormous graphs. When mapped out in a graph, wheat has 10 billion nodes of sequencing information, says Caccamo. These graphs are now considered knowledge that can be used by biologists use for the purposes of obtaining better crops.
Caccamo explains that having the right strategy in place is essential for the long term success of TGAC because their data growth is exponential. He notes that as the data becomes cheaper and cheaper to process, the scope of the research tends to expand. In the example of wheat genetics, he explains that the research has expanded to examine environmental factors.
“A recent addition of the toolkit of the bioinformatician is that we can look now into what is present as well in the soil,” says Caccamo. “In this case, it’s not going to be about one species, but instead about a community of species – what we call a microbial community. That’s what we call metagenomics.”
While the concept of metagenomics is contained within the esoteric domain of the genomic community, the concept of the run-away project is not. Virtually every enterprise has experienced the resource drain that happens when a project expands beyond its original scope. In the case of big data, these run-away project can be very costly if there aren’t strategies in place to govern the direction they take.
As enterprises ramp up their big data initiatives, TGAC’s example suggests that organizations would be wise to consider the culture and processes already in place where traditional sciences have already blazed a trail in managing and processing enormous amounts of data.
Related Items:
Boosting Big National Lab Data
Intel CIO’s Big Data Prescription
July 3, 2025
- FutureHouse Launches AI Platform to Accelerate Scientific Discovery
- KIOXIA AiSAQ Software Advances AI RAG with New Version of Vector Search Library
- NIH Highlights AI and Advanced Computing in New Data Science Strategic Plan
- UChicago Data Science Alum Transforms Baseball Passion into Career with Seattle Mariners
July 2, 2025
- Bright Data Launches AI Suite to Power Real-Time Web Access for Autonomous Agents
- Gartner Finds 45% of Organizations with High AI Maturity Sustain AI Projects for at Least 3 Years
- UF Highlights Role of Academic Data in Overcoming AI’s Looming Data Shortage
July 1, 2025
- Nexdata Presents Real-World Scalable AI Training Data Solutions at CVPR 2025
- IBM and DBmaestro Expand Partnership to Deliver Enterprise-Grade Database DevOps and Observability
- John Snow Labs Debuts Martlet.ai to Advance Compliance and Efficiency in HCC Coding
- HighByte Releases Industrial MCP Server for Agentic AI
- Qlik Releases Trust Score for AI in Qlik Talend Cloud
- Dresner Advisory Publishes 2025 Wisdom of Crowds Enterprise Performance Management Market Study
- Precisely Accelerates Location-Aware AI with Model Context Protocol
- MongoDB Announces Commitment to Achieve FedRAMP High and Impact Level 5 Authorizations
June 30, 2025
- Campfire Raises $35 Million Series A Led by Accel to Build the Next-Generation AI-Driven ERP
- Intel Xeon 6 Slashes Power Consumption for Nokia Core Network Customers
- Equal Opportunity Ventures Leads Investment in Manta AI to Redefine the Future of Data Science
- Tracer Protect for ChatGPT to Combat Rising Enterprise Brand Threats from AI Chatbots
June 27, 2025
- Inside the Chargeback System That Made Harvard’s Storage Sustainable
- What Are Reasoning Models and Why You Should Care
- Databricks Takes Top Spot in Gartner DSML Platform Report
- LinkedIn Introduces Northguard, Its Replacement for Kafka
- Change to Apache Iceberg Could Streamline Queries, Open Data
- Agentic AI Orchestration Layer Should be Independent, Dataiku CEO Says
- Why Snowflake Bought Crunchy Data
- Fine-Tuning LLM Performance: How Knowledge Graphs Can Help Avoid Missteps
- Top-Down or Bottom-Up Data Model Design: Which is Best?
- The Evolution of Time-Series Models: AI Leading a New Forecasting Era
- More Features…
- Mathematica Helps Crack Zodiac Killer’s Code
- ‘The Relational Model Always Wins,’ RelationalAI CEO Says
- Confluent Says ‘Au Revoir’ to Zookeeper with Launch of Confluent Platform 8.0
- DuckLake Makes a Splash in the Lakehouse Stack – But Can It Break Through?
- Solidigm Celebrates World’s Largest SSD with ‘122 Day’
- The Top Five Data Labeling Firms According to Everest Group
- Supabase’s $200M Raise Signals Big Ambitions
- Toloka Expands Data Labeling Service
- With $17M in Funding, DataBahn Pushes AI Agents to Reinvent the Enterprise Data Pipeline
- Databricks Is Making a Long-Term Play to Fix AI’s Biggest Constraint
- More News In Brief…
- Astronomer Unveils New Capabilities in Astro to Streamline Enterprise Data Orchestration
- Databricks Unveils Databricks One: A New Way to Bring AI to Every Corner of the Business
- Seagate Unveils IronWolf Pro 24TB Hard Drive for SMBs and Enterprises
- Gartner Predicts 40% of Generative AI Solutions Will Be Multimodal By 2027
- BigBear.ai And Palantir Announce Strategic Partnership
- Astronomer Introduces Astro Observe to Provide Unified Full-Stack Data Orchestration and Observability
- Databricks Donates Declarative Pipelines to Apache Spark Open Source Project
- Deloitte Survey Finds AI Use and Tech Investments Top Priorities for Private Companies in 2024
- Code.org, in Partnership with Amazon, Launches New AI Curriculum for Grades 8-12
- Databricks Announces Data Intelligence Platform for Communications
- More This Just In…