Architecting Your Data Lake for Flexibility

The terms ‘Big Data’ and ‘Hadoop’ have come to be almost synonymous in today’s world of business intelligence and analytics. The promise of easy access to large volumes of heterogeneous data, at low cost compared to traditional data warehousing platforms, has led many organizations to dip their toe in the water of a Hadoop data lake. The Life Sciences industry is no exception.
With the extremely large amounts of clinical and exogenous data being generated by the healthcare industry, a data lake is an attractive proposition for companies looking to mine data for new indications, optimize or accelerate trials, or gain new insights into patient and prescriber behavior.
Often, the results do not live up to their expectations. It’s one thing to gather all kinds of data together, but quite another to make sense of it. Hadoop was originally designed for relatively small numbers of very large data sets. Bringing together large numbers of smaller data sets, such as clinical trial results, presents problems for integration, and when organizations are not prepared to address these challenges, they simply give up. The data lake turns into a ‘data swamp’ of disconnected data sets, and people become disillusioned with the technology.
However, the perceived lack of success in many Hadoop implementations is often due not to shortcomings in the platform itself, but instead with users’ preconceived expectations of what Hadoop can deliver and with the way their experiences with data warehousing platforms have colored their thinking.
Too many organizations simply take their existing data warehouse environments and migrate them to Hadoop without taking the time to re-architect the implementation to properly take advantage of new technologies and other evolving paradigms such as cloud computing. A data lake is not a data warehouse, and while many of the architectural principles developed over 20+ years of data warehousing can be applied to a data lake, many others simply don’t work, at least not without some adaptation.
Some of these changes fly in the face of accepted data architecture practices and will give pause to those accustomed to implementing traditional data warehouses. The remainder of this article will explain some of the mind shifts necessary to fully exploit Hadoop in the cloud, and why they are necessary.
Big, Fast, or Cheap? Make Your Choice
We are all familiar with the four Vs of Big Data:
- Volume: store and process enormous volumes of data
- Velocity: ingest and process data promptly
- Variety: store and process diverse data types
- Veracity: ensure the data we process is clean and accurate
The core Hadoop technologies such as Hadoop Distributed File System (HDFS) and MapReduce give us the ability to address the first three of these capabilities and, with some help from ancillary technologies such as Apache Atlas or the various tools offered by the major cloud providers, Hadoop can address the veracity aspect too.
Some people have taken this to mean a Hadoop platform can deliver all of these things simultaneously and in the same implementation. This is not the case. As with any technology, some trade-offs are necessary when designing a Hadoop implementation. There are many details, of course, but these trade-offs boil down to three facets as shown below.
- Big refers to the volume of data you can handle with your environment. Hadoop allows you to scale your storage capacity – horizontally as well as vertically – to handle vast volumes of data.
- Fast refers to the speed with which you can ingest and process the data and derive insights from it. Hadoop allows you to scale your processing capacity using relatively cheap commodity hardware and massively parallel processing techniques to access and process data quickly.
- Cheap refers to the overall cost of the platform. This means not just the cost of the infrastructure to support your storage and processing requirements, but also the cost of building, maintaining and operating the environment which can grow quite complicated as more requirements come into play.
The bottom line here is that there’s no magic in Hadoop. Like any other technology, you can typically achieve one or at best two of these facets, but in the absence of an unlimited budget, you typically need to sacrifice in some way.
This is not necessarily a bad thing. It merely means you need to understand your use cases and tailor your Hadoop environment accordingly. If you want to analyze petabytes of data at relatively low cost, be prepared for those analyses to take a significant amount of processing time. If you want to analyze data quickly at low cost, take steps to reduce the corpus of data to a smaller size through preliminary data preparation. If you want to analyze large volumes of data in near real-time, be prepared to spend money on sufficient compute capacity to do so.
You’re probably thinking ‘how do I tailor my Hadoop environment to meet my use cases and requirements when I have many use cases with sometimes conflicting requirements, without going broke?
Embrace the Cloud
Cloud computing has expanded rapidly over the past few years, and all the major cloud vendors have their own Hadoop services. There’s very little reason to implement your own on-premise Hadoop solution these days, since there are few advantages and lots of limitations in terms of agility and flexibility.
Rather than investing in your own Hadoop infrastructure and having to make educated guesses about future capacity requirements, cloud infrastructure allows you to reconfigure your environment any time you need to, scale your services to meet new or changing demands, and only pay for what you use, when you use it. We’ll talk more about these benefits later.
Be a Splitter! Separate Storage from Compute Capacity
Many early adopters of Hadoop who came from the world of traditional data warehousing, and particularly that of data warehouse appliances such as Teradata, Exadata, and Netezza, fell into the trap of implementing Hadoop on relatively small clusters of powerful nodes with integrated storage and compute capabilities.
One pharma company migrated their data warehouse to Hadoop on a private cloud, on the promise of cost savings, using a fixed-size cluster that combined storage and compute capacity on the same nodes. Effectively, they took their existing architecture, changed technologies and outsourced it to the cloud, without re-architecting to exploit the capabilities of Hadoop or the cloud. Not surprisingly, they ran into problems as their data volume and velocity grew since their architecture was fundamentally at odds with the philosophy of Hadoop.
Far more flexibility and scalability can be gained by separating storage and compute capacity into physically separate tiers, connected by fast network connections. This allows you to scale your storage capacity as your data volume grows and independently scale your compute capacity to meet your processing requirements.
Let’s say you’re ingesting data from multiple clinical trials across multiple therapeutic areas into a single data lake and storing the data in its original source format. Some of the trials will be larger than others and will have generated significantly more data. You can decide how big a compute cluster you want to use, depending on how fast you want to ingest and store the data, which depends on its volume and velocity, but also on the amount of data cleansing you anticipate doing, which depends on the data’s veracity.
It’s dangerous to assume all data is clean when you receive it. There may be inconsistencies, missing attributes etc. That doesn’t mean you should discard those elements though, since the inconsistencies or omissions themselves tell you something about the data. There can often be as much information in the metadata – implicit or explicit – as in the data set itself.
Once you’ve successfully cleansed and ingested the data, you can persist the data into your data lake and tear down the compute cluster. That way, you don’t pay for compute capacity you’re not using, as described below.
To take the example further, let’s assume you have clinical trial data from multiple trials in multiple therapeutic areas, and you want to analyze that data to predict dropout rates for an upcoming trial, so you can select the optimal sites and investigators. You can use a compute cluster to extract, homogenize and write the data into a separate data set prior to analysis, but that process may involve multiple steps and include temporary data sets.
Separating storage capacity from compute capacity allows you to allocate space for this temporary data as you need it, then delete the data sets and release the space, retaining only the final data sets you will use for analysis. That means you’re only paying for storage when you need it. Again, we’ll talk about this later in the story.
Separate Ingestion, Extraction and Analysis
Separating storage from compute capacity is good, but you can get more granular for even greater flexibility by separating compute clusters. Compute capacity can be divided into several distinct types of processing:
- Ingestion loads data into the data lake, either in batches or streaming in near real-time. Ingestion can be a trivial or complicated task depending on how much cleansing and/or augmentation the data must undergo.
- Extraction takes data from the data lake and creates a new subset of the data, suitable for a specific type of analysis. The data may be further augmented, integrated or transformed from its original form.
- Analysis takes raw or transformed data and derives insights from it. This can be descriptive analysis to create reports on historical data, predictive analysis using mathematical modeling, machine learning, or other techniques, or prescriptive analysis providing reactive outputs to stimulus events within the data.
A lot of organizations fall into the trap of trying to do everything with one compute cluster, which quickly becomes overloaded as different workloads with different requirements inevitably compete for a finite set of resources. The typical response to that is to add more capacity, which adds more expense and decreases efficiency since the extra capacity is not utilized all the time.
For optimum efficiency, you should separate all these tasks and run them on different infrastructure optimized for the specific task at hand. You can stand up a cluster of compute nodes, point them at your data set, derive your results, and tear down the cluster, so you free up resources and don’t incur further cost.
In our previous example of extracting clinical trial data, you don’t need to use one compute cluster for everything. Exploring the source data sets in the data lake will determine the data’s volume and variety, and you can decide how fast you want to extract and potentially transform it for your analysis.
Those factors will determine the size of the compute cluster you want and, in conjunction with your budget, will determine the size of the cluster you decide to use. You can then use a temporary, specialized cluster with the right number and type of nodes for the task and discard that cluster after you’re done.
That extraction cluster can be completely separate from the cluster you use to do the actual analysis, since the optimal number and type of nodes will depend on the task at hand and may differ significantly between, for example, data harmonization and predictive modeling. Don’t be afraid to separate clusters. In the cloud, compute capacity is expendable. Stand up and tear down clusters as you need them.
Exploit Elastic Capacity and Pay-For-What-You-Use Charging
In the “Separate Storage from Compute Capacity” section above, we described the physical separation of storage and compute capacity. You can gain even more flexibility by leveraging elastic capabilities that scale on demand, within defined boundaries, without manual intervention.
Furthermore, elastic capacity allows you to scale down as well as upward. Storage requirements often increase temporarily as you go through multi-stage data integrations and transformations and reduce to a lower level as you discard intermediate data sets and retain only the result sets. Compute capacity requirements increase during complex integrations or analyses and drop significantly when those tasks are complete.
Using our trial site selection example above, you can discard the compute cluster you use for the modeling after you finish deriving your results. You may even want to discard the result set if the analysis is a one-off and you will have no further use for it. In this way, you pay only to store the data you actually need.
To best exploit elastic storage and compute capacity for flexibility and cost containment – which is what it’s all about – you need a pay-for-what-you-use chargeback model. While many larger organizations can implement such a model, few have done so effectively. Instead, most turn to cloud providers for elastic capacity with granular usage-based pricing.
Two Tiers, Not Three
Traditional data warehouses typically use a three-tiered architecture, as shown below:
- The Staging area contains source data in its original format and is typically used as a landing zone for source systems to push data to the warehouse.
- The Canonical Data Model is the heart of the warehouse and ideally contains a single, normalized, fully integrated and enterprise-wide representation of all the data in the warehouse – though those of us familiar with data warehousing know that this is rarely the case in reality.
- Data Marts contain subsets of the data in the Canonical Data Model, optimized for consumption in specific analyses. The data in Data Marts is often denormalized to make these analyses easier and/or more performant.
The normalized, canonical data layer was initially devised to optimize storage and therefore cost since storage was relatively expensive in the early days of data warehousing. The analytics of that period were typically descriptive and requirements were well-defined. Predictive analytics tools such as SAS typically used their own data stores independent of the data warehouse.
Normalization has become something of a dogma in the data architecture world and in its day, it certainly had benefits. However, it also has a number of drawbacks, not the least of which is it significantly transforms the data upon ingestion. This transformation carries with it a danger of altering or erasing metadata that may be implicitly contained within the data. Yet many people take offense at the suggestion that normalization should not be mandatory.
Back to our clinical trial data example, assume the original data coming from trial sites isn’t particularly complete or correct – that some sites and investigators have skipped certain attributes or even entire records. If you cleanse the data, normalize it and load it into a canonical data model, it’s quite likely that you’re going to remove these invalid records, even though they provide useful information about the investigators and sites from which they originate.
In the Data Lake world, simplify this into two tiers, as follows:
The critical difference is the data is stored in its original source format. It may be augmented with additional attributes but existing attributes are also preserved. Even dirty data remains dirty because dirt can be informative. Data is not normalized or otherwise transformed until it is required for a specific analysis.
At that time, a relevant subset of data is extracted, transformed to suit the analysis being performed and operated upon. This paradigm is often called schema-on-read, though a relational schema is only one of many types of transformation you can apply.
Of course, real-time analytics – distinct from real-time data ingestion which is something quite different – will mandate you cleanse and transform data at the time of ingestion. However, if you want to the make the data available for other, as of yet unknown analyses, it is important to persist the original data.
This two-tier architecture has a number of benefits:
- It inherently preserves the original form of the data, providing a built-in archive.
- It preserves any implicit metadata contained within the data sets, which, along with the original data, facilitates exploratory analytics where requirements are not well-defined.
- It reduces storage requirements in the data lake by eliminating the canonical layer – while storage is typically cheaper in a Big Data world, it isn’t free.
- It reduces complexity, and therefore processing time, for ingestion.
Leverage the Envelope Architectural Pattern
Where the original data must be preserved but augmented, an envelope architectural pattern is a useful technique. This pattern preserves the original attributes of a data element while allowing for the addition of attributes during ingestion.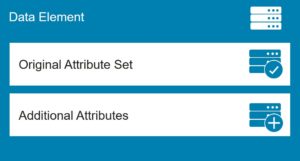
Drawing again on our clinical trial example, suppose you want to predict optimal sites for a new trial, and you want to create a geospatial visualization of the recommended sites. However, the historical data comes from multiple systems and each represents zip codes in its own way. As part of the extraction and transformation process, you can perform a look up against geospatial index data to derive the latitude and longitude coordinates for a site, and store that data as additional attributes of the data elements, while preserving the original address data.
An envelope pattern is most easily implemented in object (XML or JSON) databases but can also be implemented in any structured or semi-structured data stores such as Hive or even traditional relational database platforms.
Data Governance
We can’t talk about data lakes or data warehouses without at least mentioning data governance. Governance is an intrinsic part of the veracity aspect of Big Data and adds to the complexity and therefore to cost.
Data governance is the set of processes and technologies that ensure your data is complete, accurate and properly understood. All too many incorrect or misleading analyses can be traced back to using data that was not appropriate, which are as a result of failures of data governance.
Data governance in the Big Data world is worthy of an article (or many) in itself, so we won’t dive deep into it here. Just remember that understanding your data is critical to understanding the insights you derive from it, and the more data you have, the more challenging that becomes.
A two-tier architecture makes effective data governance even more critical, since there is no canonical data model to impose structure on the data, and therefore promote understanding. In reality, canonical data models are often insufficiently well-organized to act as a catalog for the data. Separate data catalog tools abound in the marketplace, but even these must be backed up by adequately orchestrated processes.
Summary
Many organizations have developed unreasonable expectations of Hadoop. ‘It can do anything’ is often taken to mean ‘it can do everything.’ As a result, experiences often fail to live up to expectations.
Hadoop, in its various guises, has a multitude of uses, from acting as an enterprise data warehouse to supporting advanced, exploratory analytics. However, implementing Hadoop is not merely a matter of migrating existing data warehousing concepts to a new technology. Getting the most out of your Hadoop implementation requires not only tradeoffs in terms of capability and cost but a mind shift in the way you think about data organization.
Separate storage from compute capacity, and separate ingestion, extraction and analysis into separate clusters, to maximize flexibility and gain more granular control over cost. Take advantage of elastic capacity and cost models in the cloud to further optimize costs. Reduce complexity by adopting a two-stage, rather than three-stage data lake architecture, and exploit the envelope pattern for augmentation while retaining the original source data.
If you embrace the new cloud and data lake paradigms rather than attempting to impose twentieth century thinking onto twenty-first century problems by force-fitting outsourcing and data warehousing concepts onto the new technology landscape, you position yourself to gain the most value from Hadoop.
About the author:Neil Stokes is an IT Architect and Data Architect with NTT DATA Services, a top 10 global IT services provider. With more than 30 years of experience in the IT industry, Neil leads a team of architects, data engineers and data scientists within the company’s Life Sciences vertical. For the past 15 years he has specialized in the Healthcare and Life Sciences industries, working with Payers, Providers and Life Sciences companies worldwide.
Related Items:
Rethinking Architecture at Massive Scale
Re-Imagining Big Data in a Post-Hadoop World