Accelerating Drug Discovery with Machine Learning on Big Medical Data

Pharmaceutical companies spend billions testing prospective drugs by conducting “wet lab” experiments that can take years to complete. But what if the same results could be obtained in a matter of minutes by running computer model simulations instead? A Silicon Valley startup says it has created a novel machine learning algorithm that does just that.
twoXAR (pronounced “two-czar”) was founded last year by two men both named Andrew Radin (more on that later). The Radins were interested in using advances in data science and large-scale computing to speed up the pace of drug discovery, which would give pharmaceutical companies better candidates for clinical studies.
“The traditional method of drug discovery is you examine a disease, you devise that there’s a protein of interest to you, and if you can regulate an activity with a drug, then perhaps it will help out,” twoXAR CEO Andrew A. Radin tells Datanami. “That’s a multi-year process to do all that work.”
twoXAR’s DUMA Drug Discovery platform can replace years’ worth of biological lab work with a digital equivalent that leans heavily on data science and leverages the incredible amount of medical, biological, and drug data that already exists in the public sphere.
“Our core IP [intellectual property], if you will, is this ability to take extremely diverse data sets and draw relationships between those data sets,” Radin says. “We’re combining clinical data in combination with gene expression assays, protein interaction networks, drug protein binding databases, and physical attributes about the molecules themselves.”
The hardest part is actually obtaining the relevant data. Most of the data is in the public sphere, thanks to the billions in spent by the NIH, the FDA, the European Union, and the Canadian government to back primary medical, chemical, and biological research. Once the right data is loaded into the system, it’s as simple as pressing the “go” button and waiting for the system to spit out its answer.
Digital Shortcut to Drug Discovery
The company has a patent pending for the key algorithm in DUMA, so it will remain a bit of a black box for now. But Andrew A. Radin provided Datanami a general description of how it works.
“What it does is it has a method for extracting the most relevant data–there’s some work to extract signal from noisy data from these very diverse data sets,” he says. “We then load it into a relationship model, where relationships between these data sets are calculated in the memory of the computer. And then from there we use that to create feature vectors that ultimately get loaded into a machine learning algorithm.
“At the end of the day, what we’re doing is seeing if we see the same evidence in multiple data sources,” he continues. “And if we see the same evidence coming from multiple, very diverse data sets, that gives us power to making a prediction about efficacy for a drug.”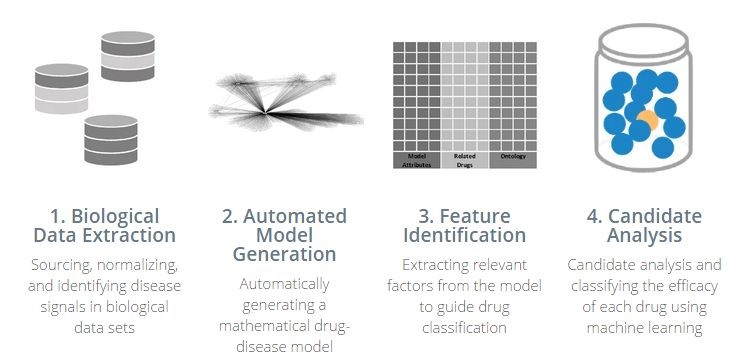
There’s a lot more to it than that, obviously. As a data scientist who has designed large-scale computer systems for some well-known companies, Andrew A. Radin understands the challenges that can trip up the most carefully planned digital experiments. At twoXAR, a big focus is following false positives down the drug-discovery rabbit hole.
“Let’s say you’re looking at clinical data for disease signal for Parkinson’s,” he says. “If I look at birth control medicine for example, that’s a drug taken by young women, and Parkinson’s is a disease for old men. If I look at birth control medicine, I might get a false signal that it’s protective against Parkinson’s.”
Gene expression micro arrays, he says, are notorious for giving you false signals. That’s why it’s so critical for twoXAR to examine other types of data, and cross-checking the results.
“When we see the evidence, if you will, from these different data sources sort of pile up…that brings confidence to an the ultimate prediction that you’re making–that it’s not because of a confounding error or a false discovery rate error, but that there’s something meaningful to make a prediction, because it’s repeated in multiple data sets,” he says.
Feedback Loops
The company checks DUMA’s accuracy by feeding data about known, FDA-approved treatments into the machine (which runs on the AWS cloud) and seeing what it spits out. “We’ll remove the label so the algorithm itself doesn’t know that this is a known treatment, and run this through the prediction engine,” Andrew A. Radin says.
“So you’d expect with type 2 diabetes, for a drug like Metformin darn well better come out the other end as highly predictive of the disease. If they don’t…that tells us that we need to go out and collect more data or improve the diversity of the data to improve the quality of the prediction.”
While the DUMA Drug Discovery system can work with any type of disease, so far, the young company has focused its efforts primarily on cardiovascular, metabolic, and neurological diseases. The company is currently working with university researchers and major pharmaceutical companies to prove that its system works.
This type of computational drug discovery is new to the major drug companies, so it’s going to take some time and effort to prove that approaches like twoXAR’s are valid. That means doing wet lab work that duplicates and validates the output of the DUMA Drug Discovery system.
“All the pharmaceutical companies we’ve interacted with have all said the same thing. ‘We don’t know how to evaluate the computational methodologies, but we do know how to evaluate results of a pre- clinical study. Take your predictions, test those predictions in an animal model, and then we’re ready to do partnerships with you,'” Radin says.
Early Success
twoXAR has done a bit of work with the NIH Udall Center of Excellence in Parkinson’s Disease Research at Michigan State University. In fact, the company’s first trip to the lab had an interesting end.
“We loaded a bunch of data on Parkinson’s Disease into the system, pressed the go button, a few minutes later we had a list of drugs that were listed as highly efficacious,” Andrew A. Radin says. “So we started Googling the results, and one of the top hits in our prediction was a drug [that we saw] had been under study in the NIH Parkinson’s research lab at Michigan State that Dr. Tim Collier had been working on it for years.”
The twoXAR employees decided to blindly email him and offer up the computational evidence for the drug. The twoXAR executive didn’t expect to hear back, but the next day, Dr. Collier called him on the phone.
“He called and said ‘You guys just came up with something that backed up what I’ve been doing in the lab for years,” he says. “He said, would you do another of these drugs. So we ran it through the system, and handed the results back to him. Now a bunch of people in his lab were all excited and the reason why is we have, sort of instantaneously if you will, produced evidence that matches their understanding about what’s going on in this particular drug and why it’s a potential good treatment for Parkinson’s.”
twoXAR is currently working with the NIH Parkinson’s lab at Michigan State to validate the lab’s work with the computational model. The lab is also running pre-clinical animal studies to validate new drug candidates that were identified by DUMA. It’s also working with several pharmaceutical companies on proof of concepts (POCs).
Who Is Andrew Radin?

Andrew M. Radin (left), co-founder and Chief Business Officer, and Andrew A. Radin, co-founder and CEO of twoXAR
A serendipitous meeting of the two Andrew Radins put the two men on course for the founding of twoXAR (which can be read as “2x Andrew Radin.”). As the co-founders write in their joint blog at www.andrewradin.com, the original meeting had to do with that very domain name.
Several years back, Andrew A. Radin was surprised when Andrew M. Radin wanted to buy the domain that bears their name. Andrew A. Radin told him in no uncertain terms that he was not interested. Eventually, the contentious emails died down, common interests emerged, and a real friendship resulted.
At some point, the two Andrew Radins decided to go into business together that capitalized on their joint interested and aptitude for technology and science. Andrew A. Radin had followed up his tech startup days by studying bioinformatics at Stanford. Andrew M. Radin, meanwhile, had attended MIT’s Sloan School of Management and worked as an investor in venture and private equity funds.
Related Items:
Creating Flexible Big Data Solutions for Drug Discovery
Five Ways Big Genomic Data Is Making Us Healthier
Big Data’s Next Big Thing: Sports Training and Personalized Medicine