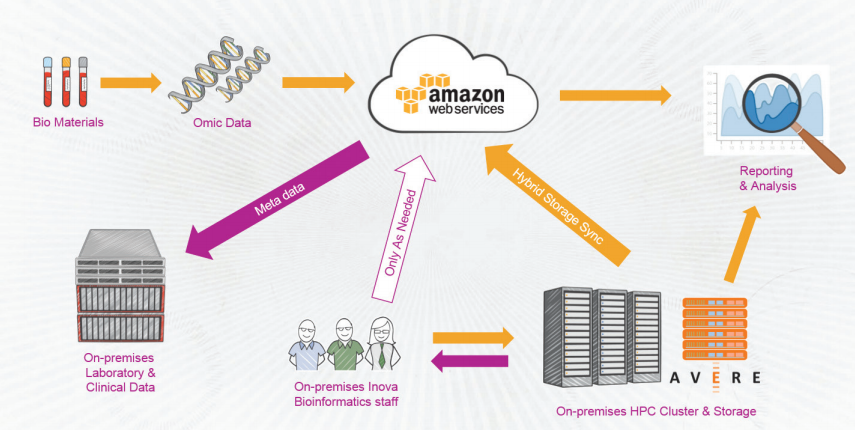
Peek Inside a Hybrid Cloud for Big Genomics Data
Genomics, or the study of human genes, holds an enormous promise to improve the health of people. However, at 80GB to 250GB per sequenced genome, the sheer size of genomics data poses a considerable storage and processing challenge. To keep costs in check, Inova Translational Medical Institute (ITMI) adopted a hybrid cloud architecture for its ongoing genomics studies.
ITMI is a branch of the non-profit Inova hospital network in the Washington D.C. area that received 150 million in funding by Inova to pursue the field of genomics. The Falls Church, Virginia organization has launched six separate studies in the fields of genomics, including a premature birth study, a Type 2 diabetes study, and a congenital disorders study, among others.
When ITMI received its grant in 2011, the institute was not sure where the research would take it, so it opted for practical and simple approach: It chose to store genomics data on the Amazon Simple Storage Service (S3) public cloud rather than keep it on premise. To get gene sequence data into S3, it would ship hundreds of hard disks back and forth across the country to and from its gene sequencing partner, Illumina, based 3,000 miles away in San Diego.
After receiving gene sequence data from Illumina, the ITMI analysts would check the data for errors, and then upload it to S3, which itself was an error-prone process that could take up to a week to complete. This workflow obviously wasn’t ideal, but it was relatively affordable. Besides, when ITMI started, it had the gene sequences of several hundred people to study–representing perhaps 50 terabytes or so—so the size of the data wasn’t a major issue.
The Genomic Growth Curve
But over the years, as ITMI launched more studies, the size and variety of the data has exploded and become a stumbling block. Today, ITMI maintains close to 7,200 human genomes, representing 1.7PB of genomic data stored on Amazon’s S3 cloud, according to Aaron Black, director of informatics ITMI. And with plans to expand the genomic studies to 20,000 people by 2017—or eight to 10 genomes uploaded per day–there’s no end in sight for the big data crunch.
However, Amazon’s cloud-based object storage system is just one leg of a three-legged data stool that ITMI relies on to do its research. In addition to S3, the institute access clinical data and diagnoses stored in an Epic electronic medical record (EMR) system that Inova spend hundreds of millions of dollars to build. The third leg, which went online just a year ago, is a 1,024-core UV2000 supercomputer from SGI that’s used to crunch genomics data.
At about the time that SGI high performance computing (HPC) system went live, ITMI decided to reevaluate its data architecture—in particular whether it was time to bring the Amazon S3 data back in house.
“We did a cost analysis of what it would look like in the next three years,” Black says. “Based on what we were paying at Amazon and what they thought they would have to build or expand in their current data center, it was tens of millions of dollars difference. Amazon was so much cheaper.”
While Amazon S3 is saving ITMI tens of millions, it still lacked that “glue” that could bring these three elements together while maintaining the necessary security and privacy controls. It ended up selecting a NAS storage appliance from Avere Systems that could blend the three data sources together into a seamless whole.
Splicing Hybrid Cloudiness
“What Avere can do is make sure the Amazon buckets…look like they’re locally mounted so researchers can see the data,” Black says. ITMI works with researchers around the world and Avere provides fine-grained control over which data sets the researchers can access. The institute also uses the encryption key management software provided by the Avere edge filers.
Being located just a couple of miles from the Amazon data center is a nice benefit for ITMI, and so is the big 10Gbps pipe linking them. But even with all that bandwidth and 1.4 PB of storage attached to the SGI super, ITMI must be careful about which data sets it chooses to load onto its HPC system. That’s where another Avere feature comes in handy.
“Avere has a special AMI [Amazon Machine Image] where they can synch certain kinds of data that we know are going to be used, so we can save it down on the ground,” Black says. “As we use it, Avere has a custom algorithm that can tell us what’s hot and not and move it into the fastest caching area.”
This helps researchers narrow their search considerably before launching jobs on the SGI cluster. “If there are 7,000 individuals, they might only want to look at kids with asthma and that might only be 70 genomes,” Black says. “They have to find a way to broker, or find those individual objects on Amazon, and then they might want to know just a subset of genes.”
Battling Data Bottlenecks
But bottlenecks still emerge, even with Avere serving as the traffic cop for data moving among the SGI cluster, Amazon S3, and the Epic EMR system. “We work with Amazon almost weekly to understand how can we get the best performance out of this,” Black says. “And what we were seeing was, it was really slow to move up. The bottleneck wasn’t the pipe, it wasn’t Avere–it was writing to disk at Amazon.”
ITMI has done a couple of things to address that issue, including adopting hashing algorithms on Amazon to make the disk IOPS more parallel and get closer to filling up the pipe. “Not that it’s horrible, but we’re spending a lot of money on a high bandwidth connection and we expect the best performance,” Black says.
The wait times are not a big issue today for ITMI’s researchers; they’re used to waiting for answers during long periods of unsupervised learning. But as ITMI ramps up its clinical practice, it will keep much “hotter” data storage and processing capacity in place, such as when it’s called on to determine what genetic condition may be impacting a newborn in the neonatal intensive care unit (NICU).
“In a clinical setting, we’d do all the computing and analysis on the HPC, on the SGI,” Black says. “Once we did the analysis, we’d move [the raw data] up to Amazon, and have a vault on the East Coast and one on West Coast, because we’d want to span zones.”
ITMI’s journey into genomics is just getting started, but it’s already showing promise. The congenital anomalies study, for example, boasts a 60 percent success rate. There’s no question genomics poses great promise to help human health. The question for the system and data architects, then, becomes how best to build a system that can scale while delivering the necessary performance.
“This is the first time we’ve been around a hospital system that’s done this at scale,” Black says. “If look at every single component of it–from storage to compute to how big your pipes have to be to what’s your long term storage strategy–every one of those components has to be analyzed. Cloud scales very well for us, form a storage perceptive. From a compute perspective, we’re not there yet [with cloud].”
Related Items:
Google Targets Big Genome Data
Data Tools Help Researchers Tackle Pediatric Cancer
Detecting Cancerous RNA Signatures With Big Data