MapReduce Alternatives in Bioinformatics
Over the last decade, MapReduce has emerged as a popular software framework for processing big data sets on large clusters of commodity hardware. The technique came out of a paper published by Google in 2004, which explains: “Users specify a map function that processes a key/value pair to generate a set of intermediate key/value pairs, and a reduce function that merges all intermediate values associated with the same intermediate key. Many real world tasks are expressible in this model…”
Accelerated by the 21st century data explosion, MapReduce quickly gained momentum in both industry and academic circles. Tools based on MapReduce have been quite useful for applications in a number of science disciplines, including bioinformatics, ecology and geoinformatics.
Distributed data-parallelization (DDP) patterns, like MapReduce, serve as reusable practices for efficient design and execution of big data analysis and analytics applications. In the last few years, other DDP patterns have been proposed, which like MapReduce combine data partition, parallel computing and distributed computing elements.
A team of computer scientists have identified viable alternatives to MapReduce for implementing big data analysis tools. In their paper “Comparison of Distributed Data-Parallelization Patterns for Big Data Analysis: A Bioinformatics Case Study,” researchers Jianwu Wang, Daniel Crawl, and Ilkay Altintas (from San Diego Supercomputer Center) together with Kostas Tzoumas and Volker Markl (from the Technische Universität Berlin) explore other DDP options.
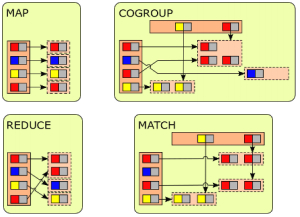 The experiment implements an existing bioinformatics tool, called CloudBurst, using three different DDP pattern combinations: MapReduce, MapCoGroup and MapMatch. The team’s goal is first, to assess whether changing the DDP pattern would have a significant effect on performance (measured in execution time), and second to identify the key factors influencing performance outcomes.
The experiment implements an existing bioinformatics tool, called CloudBurst, using three different DDP pattern combinations: MapReduce, MapCoGroup and MapMatch. The team’s goal is first, to assess whether changing the DDP pattern would have a significant effect on performance (measured in execution time), and second to identify the key factors influencing performance outcomes.
CloudBurst is a parallel seed-and-extend sequence mapping tool originally based on MapReduce. The algorithm finds sub-strings called seeds with exact matches in both the reads, or query sequences, and the reference sequences, and then extends the seeds into longer, inexact matches. The length of a seed is decided by parameter mismatches (k) and another parameter called reads length (m) with formula m/(k+1).
The team hypothesized that any significant performance differences would be due to two main factors:
- The difference between the numbers of keys in the two input data, denoted as p.
- The average number of values per query/reference key, denoted as q.
“Total execution time of Reduce/CoGroup/Match includes two main parts: user function execution time and overhead to load the user function and input data,” they write. “By looking into how Reduce pattern and Match pattern work, we identify two main factors that could affect their performances. The first factor is the difference between the numbers of keys in the two input data, denoted as p. The second factor is the average number of values per query/reference key, denoted as q.”
For the DDP execution engine, the team chose Stratosphere (version 0.2) because it supports Map, Reduce, CoGroup and Match. The experimental test bed consisted of five compute nodes in a compute cluster environment, each node equipped with two four-core CPUs.
The first experiment employed two large datasets: a query dataset of over nine million sequences from the 1000 Genomes Project and a reference dataset of over 1.2 million sequences from the CAMERA project. The resulting execution times for all three DDP implementations of CloudBurst increased when k value (the similarity threshold) increased. For the different DDP implementations, the performance of MapMatch is better than MapReduce for k=0-2, but worse when k=3. MapCoGroup’s execution times remain in between those of MapReduce and MapMatch in all cases.
 The second experiment processes only a large reference data set. The reference dataset is the same as in the first round and the query dataset only includes the first 5,000 sequences of the query dataset from the 1,000 Genomes Project used in the last experiment. Here, the execution times show MapMatch outperforming MapReduce in all cases. MapCoGroup performs second-best, and MapReduce performs worst.
The second experiment processes only a large reference data set. The reference dataset is the same as in the first round and the query dataset only includes the first 5,000 sequences of the query dataset from the 1,000 Genomes Project used in the last experiment. Here, the execution times show MapMatch outperforming MapReduce in all cases. MapCoGroup performs second-best, and MapReduce performs worst.
In their analysis, the authors conclude that different DDP patterns significantly influenced the execution performance of CloudBurst. A simple DDP pattern switch was able to speed up performance by almost two times in the first experiment (see Table 2). They further note that while selection can affect performance on a given tool, however no DDP pattern combination is always preferable.
“We find that although MapReduce can be used for wider range of applications with either one or two input datasets, it is not always the best choice in terms of application complexity and performance,” they write. “To understand the differences, we identified two affecting factors, namely input data balancing and value sparseness, on their performance differences.”
The results suggest MapReduce is more suitable for balanced input datasets and MapMatch is more suitable for imbalanced ones, and also that MapReduce is more suitable for condensed values per key and MapMatch is more suitable for sparse values per key.
The authors note that bioinformatics and other domains that have a need to match two input datasets could benefit from these findings. In the future, they plan to investigate other tools that are suitable for multiple DDP patterns and see how they perform on other DDP engines such as Hadoop, which will have wider applicability. They would also like to develop a way to exploit the identified factors to automatically select the best DDP pattern combination from multiple available ones.
Related Items:
UK Makes £7 3 Million Big Data Investment