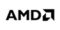Picking the Connectome Data Lock
Back in 2005, researchers at Indiana University and Lausanne University simultaneously (yet independently) spawned a concept and pet term that would become the hot topic in neuroscience for the next several years—connectomics.
The concept itself isn’t necessarily new, even thought the use of “connectomics” in popular science circles is relatively so. The idea was formed in the late 1980s and has since evolved as described succinctly below:
A hybrid between the study of genomics (the biological blueprint) and neural networks (the “connect”) this term quickly caught on, including with large organizations like the National Institutes of Health (NIH) and its Human Connectome Project.
For instance, the NIH is in the midst of a five-year effort (starting in 2009) to map the neural pathways that underlie human brain function. The purpose is to acquire and share data about the structural and functional connectivity of the human brain to advance imaging and analysis capabilities and make strides in understanding brain circuitry and associated disorders.
And talk about data… just to reconstruct the neural and synaptic connections in a mouse retina and primary visual cortex involved a 12 TB data set (which incidentally is now available to all at the Open Connectome Project).
Mapping the connectome requires a complete mapping process of the neural systems on a neuron-by-neuron basis, a task that requires accounting for billions of neurons, at least for most larger, complex mammals. According to Open Connectome Project, the human cerebral cortex alone contains something in the neighborhood of 1010 neurons linked by 1014 synaptic connections.
That number is a bit difficult to digest without context, so how about this: the number of base-pairs in a human genome is 109.
NEXT — The Big Data Challenges of Connectomics >>>
The Big Data Challenges of Connectomics
To build out the human connectome at the microscale level required (there are already some macroscale efforts based on MRI), there are serious data collection, machine vision tool-related and algorithmic hurdles that must be addressed.
While there are groups working on new innovations in serial electron microscopes to move ahead on the machine-vision and image-prcocessing side, the sophisticated pattern recognition software tools are still lagging behind.

According to Sebastian Seung, Professor of Computational Neuroscience at MIT and author of the book, Connectome, if we can collectively tap the brainpower of the planet (that’s a total of over 7 billion of us) we might be able to fully understand the brain as a singular unit.
To put the Connectome concept into some kind of context, Seung asks us to envision the crisscrossing network of airline networks in a map. He says that if each city on that map was a neuron, and every flight between two cities was a connection, it would create a diagram that goes pretty far in terms of showing a neuron network.
But in terms of the brain, that network expands to mind-boggling proportions. Now imagine that expanded to 100 billion cities and ten thousand flights from every city. That would be the complexity of the map of the network inside our brains.
Seung says that the value of the Connectome concept can be traced back to an old old hypothesis in neuroscience that suggests our memories are somehow encoded in that crisscrossing network diagram—and that each Connectome is unique. “It may be that your particular pattern of connections has a lot of information inside that huge wiring diagram. That might actually contain the information of your memories.”
He suggests that memories create the idiosyncratic aspects of our personal identities. Further, maybe your personality, maybe even people who have problems with psychiatric disorders that might also be due to some kind of special different kind of pattern in brain wiring.
To this end he claims that while his genome might not relate anything about individuality, a Connectome certainly will—but that individuality might change over time, just as our personalities do.
As Seung says, “That map of your brain s not fixed, but your experiences can alter it. That’s one reason why we believe that experiences are leaving the memories inside your brain through the Connectome.”
The question is, how can scientists tap the massive data required to reverse engineer the Connectome—to get to the heart of what makes us individuals? This is a delicate, intricate machine, he says. “You’ve got a machine, you want to take it apart, dissasamble it,” he says. “Those of you out there who are real super nerds, super engineers, will resonate with the idea of disassembling a machine to figure out how it works.”
 The difference here is that scientists can’t reverse engineer using the rip and read method. It’s impossible (not to mention gross) to think about pulling the brain apart to hone in on the individual neuron level and even if it was possible, the networks are tightly, impossibly tangled… part of what makes the Connectome idea so interesting in the first place.
The difference here is that scientists can’t reverse engineer using the rip and read method. It’s impossible (not to mention gross) to think about pulling the brain apart to hone in on the individual neuron level and even if it was possible, the networks are tightly, impossibly tangled… part of what makes the Connectome idea so interesting in the first place.
Seung has a radical approach to reverse engineering these problems. His team embeds a (dead) brain in really hard plastic resins and slice it into extremely thin slices–a thousand times thinner than a hair. They then image these brain slices in an electron microscope, which offers extremely high resolution.
What this creates is the closest thing possible to a virtual brain. It offers an image of every neuron and every synapse inside each slice. In order to map out that piece of brain, however, researchers have to trace the trajectory of every branch of a neuron through those images and find those synapses. That turns out to be a very time consuming task—and an exercise in some seriously data-intensive computing.
NEXT — Inside the Brain Slice Data…>>>
Inside the Brain Slice Data
The images from one cubic millimeter of brain, amount to about a petabyte of data if you image it at that resolution.
 As Seung says, “I want to emphasize that I’m talking about really high resolution microscopy here, not an MRI scan like you see in the newspapers. Their cubic millimeter would be one pixel, but here we’re talking about a petapixel.
As Seung says, “I want to emphasize that I’m talking about really high resolution microscopy here, not an MRI scan like you see in the newspapers. Their cubic millimeter would be one pixel, but here we’re talking about a petapixel.
That data is so overwhelming, that we are still struggling to deal with that kind of deluge. One way is to use AI, artificial intelligence to see the paths in those images. But AI still isn’t perfect, so we need people to interact with the AI, to guide it and to correct it.”
Seung’s team is inviting the public to join in the brain slice-inspired AI fun via a new site called eyewire.org.
There, potential contributors to the project are faced with the image of the retina, the neural tissue at the back of the eye. It serves that up on the web in your web browser and you can join an online community and help the team map out the neural connections. I think it’s an exciting way of getting everyone involved in a kind of endeavor that formally only professionals could be involved in.
“It’s a game and it’s a game that we’ve all enjoyed at one time in our lives. It’s like a gigantic three-dimensional coloring book. You just have to stay between the lines and color in the branch of a neuron as you follow it through this three-dimensional stack of images. It’s at its preliminary stages.
The first result is that it can be fun. We have addicts, we don’t have a lot of users yet, but we have some people who are very dedicated and excited. We have a sense of community, if you go to the discussion forums, you’ll see all kinds of great comments, and questions, and hilarious things that people think of, creative things, things that enhanced our site. We’ve learned a lot from our users.”
Related Stories
7 Big Winners in the U.S. Big Data Drive
Inside Europe’s “Super Data Cluster”
Data Miners Excavate Human Brain